International Journal of Oral and Craniofacial Science
Prenatal Alcohol Exposure Alters Osteoblast Gene Expression in Newborn Rats – Screening Study
1Department of Biosciences and Oral Diagnosis, Institute of Science and Technology, UNESP – São Paulo State University, 777 Eng. Francisco José Longo Ave. São José dos Campos, SP 12245-000, Brazil
2Institute of Research and Development – IP&D, Dynamics of Cellular Compartments Laboratory, UNIVAP - University of Vale do Paraíba. 2911 Shishima Hifumi Ave. São José dos Campos, SP 12244-390, Brazil
3Department of Science and Technology, Institute of Science and Technology, Federal University of São Paulo, 330 Talim Street, São José dos Campos, SP 12231-280, Brazil
4Multicampi School of Medical Sciences, Federal University of Rio Grande do Norte (UFRN), Caicó, RN, Brazil
5Department of Periodontics and Oral Medicine, University of Michigan School of Dentistry, 1011 N. University Ave. Ann Arbor, MI 48109-1078, USA
Author and article information
Cite this as
Carvalho ICS, et al. Prenatal Alcohol Exposure Alters Osteoblast Gene Expression in Newborn Rats – Screening Study. Int J Oral Craniofac Sci. 2026; 12(1): 1-15. Available from: 10.17352/ijocs.000067
Copyright License
© 2026 Carvalho ICS, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.This study examined the impact of prenatal alcohol exposure on osteoblast gene expression in newborn rats. Sixteen Wistar rats were divided into three groups based on their diet: Alcohol, Pair-fed, and Control. Each group received the specific diet for eight weeks before breeding and during the three weeks of gestation. The newborns were euthanized on the fifth day of life, and their calvaria were processed to isolate primary osteoblasts by sequential enzymatic digestion and then cultured for 10 days. After 10 days of osteogenic cell growth, the cells were processed to extract total RNA. Osteogenesis RT Profiler Rat PCR array was used to evaluate gene expression patterns among all groups. The Control group consumed more solids and liquids than the other groups (p < 0.0001). The Control (p > 0.005) and Alcohol (p > 0.005) groups showed a tendency to weight gain, while the Pair-fed group showed a tendency to lose weight (p > 0.005). Out of 84 genes analyzed, 17 showed significant differences between the Alcohol and Pair-fed groups (p < 0.05). Of these, some were downregulated, and others upregulated in the Alcohol group compared to the Pair-fed group. In conclusion, prenatal alcohol exposure alters gene expression related to osteogenic differentiation in newborn rats, with a possible consequence on fetal skeletal development.
The teratogenic effects of alcohol are well-documented, and the term “fetal alcohol spectrum disorders (FASD)” has been used to include diverse phenotypes after prenatal alcohol exposure [1-6]. Among all changes that prenatal ethanol exposure causes to the fetus and/or newborns, bone growth and quality disturbances have been studied in experimental models [7-17]. Additionally, experimental results have demonstrated an effective action of prenatal alcohol exposure on the bone tissue of the fetus and/or newborns, however, the mechanisms in which the alcohol acts on bone tissue of the newborns causing growth delay are still unclear. Under this perspective, our research group studied the effect of prenatal alcohol exposure on newborns osteoblasts. The first study demonstrated that prenatal alcohol exposure increased osteoblasts proliferation early and increased alkaline phosphatase (Alpl) activity and bone matrix formation in more advanced periods [18]. The second study demonstrated that the prenatal exposure to ethanol induced DNA damage in osteoblasts, as shown by micronucleus formation and a higher percentage of DNA in the comet tail [19]. Taken together, these results suggest that prenatal ethanol consumption has a direct effect on fetal osteoblasts.
Therefore, a gap in the literature regarding how prenatal alcohol exposure affects bone tissue in fetuses/newborns exists. Studies showed that alcohol acts on bone tissue causing various changes such as delayed ossification, body weight loss, reduced individual bone length, and ensuing overall bone growth delay [7,8,10-13,15,20-25]. However, these are consequences of the alcohol action on bone tissue, but the mechanisms by which these changes occur are still unknown. Our previous studies indicated a pathway in which alcohol affects the bone tissue of fetuses/newborns from alcoholic mothers by affecting their offspring osteoblasts [18,19[, thus, this study hypothesizes that prenatal alcohol exposure may lead to defects in bone by affecting osteoblast gene expression.
Materials and methods
Animals
All animal experiments were approved by the Ethical Committee in Research of São José dos Campos Institute of Science and Technology, UNESP – São Paulo State University (Protocol No. 01/2012-PA/CEP).
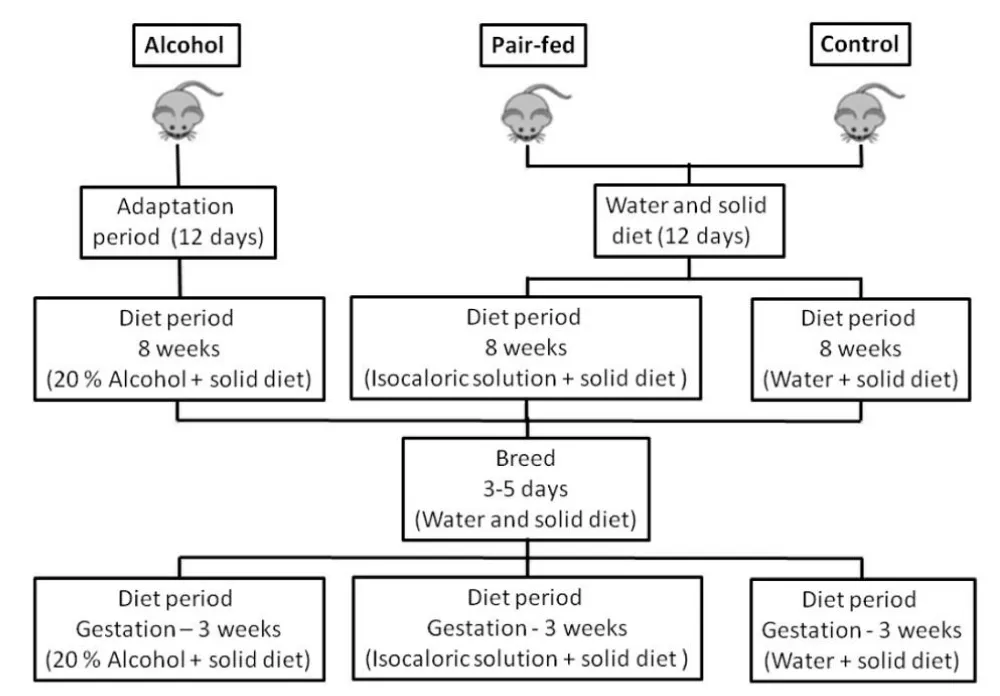
A group of sixteen nulliparous female Wistar rats, aged five weeks old, were randomly divided into three groups: i) Alcohol group, the animals were fed a 20% ethanol solution and a solid diet ad libitum. The 20% alcohol solution was obtained by dilution of absolute ethanol (ethyl alcohol P.A. ACS-99.5%, Merck, Darmstadt, Germany) in water; ii) Pair-fed group, the animals received an equivalent amount of solid diet and carbohydrate solution as the Alcohol group, and their calorie intake was matched accordingly. To achieve the calorie equivalence, the amount of alcohol and solid diet consumed by the Alcohol group was measured on the previous day, and on the following day, the Pair-fed group was given the same amount of solid diet and carbohydrate solution, as previously described [26,27], along with unrestricted access to water. iii) Control group, the animals had unrestricted access to water and solid food. The intervention involved the voluntary consumption of liquid and solid diets through oral self-administration.
After 8-weeks of dietary control, female rats were mated with Wistar males. The presence of a vaginal plug was used as an indicator of successful mating and designated as gestational day zero. The offspring remained with their mothers, who continued the same dietary regimen until euthanasia, thereby ensuring that the nutritional protocols for each group were maintained throughout the entire experimental period. At five days old, all offspring were euthanized, and their calvariae were collected for analysis.
The experimental feeding regime (Figure 1) was performed according to our previous study [19]. The female rats’ weights were assessed twice: at the beginning of the eight-week treatment protocol and after offspring euthanasia. Weight changes were calculated as a percentage using the following formula:
Blood alcohol concentration
Blood alcohol concentration in all dams was measured according to the manufacturer’s recommendations (EnzyChrom Ethanol Assay Kit, Bioassay Systems, Hayward, CA, USA) as previously described [19]. Peak blood ethanol concentrations were determined in all dams from the Alcohol group on the same day the offspring were euthanized (postnatal day 5). Following euthanasia, the dams were anesthetized for blood collection via cardiac puncture. The anesthetic agents used, along with their respective concentrations, were: xylazine (2.3 g/100 ml AnasedanV R- Vetbrands, Jacarei- Brazil), a sedative and muscle relaxant, and ketamine (1.16 g/10 ml; DopalenV R- Vetbrands, Jacarei- Brazil), a general anesthetic. An anesthetic solution was prepared containing 0.8 ml of xylazine and 0.5 ml of ketamine hydrochloride. Each animal received general anesthesia via intramuscular injection at a dose of 0.1 ml/100 g of body weight. Blood samples were collected between 8:00 and 10:00 a.m., a postprandial period corresponding to the expected peak in blood ethanol concentrations in the dams [28]. Maternal blood was centrifuged at 4°C, and the resulting serum was stored at −80 °C until ethanol concentration analysis, performed using a commercial kit. Blood samples were also collected from control dams to ensure consistent handling across groups and were used as negative controls for ethanol concentration analysis.
Cell isolation and primary culture of osteogenic cells
Primary osteoblasts were obtained from calvariae as previously described [18,19,29-40]. In brief, calvariae from 5-day-old Wistar rats were dissected and wiped with gauze. The regions containing cranial sutures were discarded, and the remaining bony tissue was cut into small pieces for digestion with a mixture of enzymes consisting of 0.2% collagenase (Thermo Fisher Scientific, Gibco, Waltham, Massachusetts, EUA) and 0.25% trypsin (Thermo Fisher Scientific, Gibco, Waltham, Massachusetts, EUA). Sequential digestion was carried out at 37°C in an agitating water bath for consecutive periods of 5, 15, and 25 minutes. The digests were centrifuged and resuspended in complete media containing α-Minimum Essential Medium with L-glutamine (Thermo Fisher Scientific, Gibco, Waltham, Massachusetts, EUA) supplemented with 10% fetal bovine serum (Thermo Fisher Scientific, Gibco, Waltham, Massachusetts, EUA), 7 mM β-glycerophosphate (Sigma/Aldrich, St. Louis, MO, EUA), 5 μg/ml ascorbic acid (Mallinckrodt Chemicals, UK), and 50 μg/ml gentamicin (Thermo Fisher Scientific, Gibco, Waltham, Massachusetts, EUA). Supernatant from the first digestion was discarded, and the second and third digests were pooled and resuspended in complete media. The pooled material was passed through a 200 µm metal screen. An aliquot of the filtrate was stained with trypan blue, and the total cell count and viability were evaluated using a hemocytometer.
Aliquots containing 1 × 10^6 cells from each experimental group were frozen at -80 °C for future use. The cells were thawed and cultured for 10 days in complete media at 37ºC in a humidified atmosphere with 5% CO2. The culture medium was replaced every three days, and the advancement of the cultures was assessed using phase contrast microscopy (Inverted Microscope OLYMPUS CK40).
For each experimental group, calvariae were collected from multiple rat pups originating from different dams. Specifically, the Control group consisted of 62 pups from 5 dams, the Pair-fed group comprised 61 pups from 5 dams, and the Alcohol group included 45 pups from 6 dams. Accordingly, the number of calvariae used for primary osteoblast isolation differed across experimental conditions. The samples were then pooled to generate a cell culture, RNA was extracted, and used for gene expression analysis by real-time PCR, following the protocol provided with the QIAGEN PCR array kit in triplicate.
Real-time PCR
After 10 days of cell culture, RNA was extracted with the RNAeasy mini kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions. The extracted RNA was quantified (ng/µL) in a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Gibco, Waltham, Massachusetts, EUA). Only RNAs presenting purity between 1.8 and 2.0 were included in this study and treated with DNase. For removal of contaminating DNA, the extracted total RNA was treated with RNase-Free DNase Set (Qiagen, Hilden, Germany), and the RNA was reverse transcribed to complementary DNA (cDNA) using the RT2 First Strand Kit (Qiagen, Hilden, Germany) according to manufacturer instructions.
Quantitative real-time PCR (qRT-PCR) was used to assess alterations in gene expression, and the StepOnePlus™ Real-Time PCR System (Applied Biosystems, Framingham, MA, USA) was employed for this purpose using a gene plate “RT2 Profiler Rat Osteogenesis – PARN 026ZA_12 as instructed by the manufacturer (Qiagen, Hilden, Germany). This plate model is composed of 84 target genes, five housekeeping genes (ACTB – l -beta muscle actin, HPRT1 - Hypoxanthine phosphoribosyl transferase 1, B2M - Beta-2-microglobulin, LDHA - Lactate dehydrogenase A and RPLP1 - Protein ribosomal greater P1), rat genomic DNA contamination control (RGDC), three controls for reverse transcription (RTC), and three positive controls (PPC) (Table 1 – Supplementary Material).
The conditions of the PCR were an initial cycle at 50 ºC for 2 min, 95 ºC for 2 min, followed by 40 cycles of 15 s at 95 ºC and 30 s at 68 ºC, and a final cycle at 72 ºC for 10 min. Upon completion of the qRT-PCR, the baseline and CT values were automatically set for each plate, and the same thresholds for all PCR array plates were kept. The data was exported and analyzed by RT2 Profiler PCR Array Data Analysis template v 3.5 (Data Analysis Center | Genetic Analysis Tools | GeneGlobe), where Fold-Change (2^ (- Delta Delta Ct)) is the result of normalized gene expression (2^(- Delta Ct)) in the Test Sample divided the normalized gene expression (2^(- Delta Ct)) in the Control Sample [41]. Fold-Regulation represents fold-change results in a biologically meaningful way. Fold-change values greater than one indicate a positive or an up-regulation, and the fold-change is equal to the fold-regulation. Fold-change values less than one indicate a negative or down-regulation, and the fold-change is the negative inverse of the fold-change.
Statistical analysis
The diet data obtained from rats exhibited a non-normal distribution and were analyzed using non-parametric tests (Kruskal-Wallis test followed by Dunn’s test). Data obtained from the weight of the rats showed normal distribution and were evaluated using parametric tests (analysis of variance –ANOVA followed by Tukey test).
The qPCR data were analyzed using software available on the company website Qiagen (Data Analysis Center | Genetic Analysis Tools | GeneGlobe). Statistical analysis of gene expression data was performed with the nonpaired Student’s t-test and considered statistically significant with p-values <0.05. Value of the fold-change greater than 2 was defined as increased expression, and the fold-change value of less than 0.5 as decreased gene expression. The analyses and comparisons were performed between the Pair-fed and Control groups, and the Alcohol and Pair-fed groups. These comparisons were made to assess whether the Pair-fed group, our Control group, showed differences in gene expression compared to the Alcohol group.
Results
Dam diet and weight
The values of solid diet, liquid diet, and the changes in the weight (percentage) were shown in Table 1 (changes in body weight were calculated based on measurements obtained at the beginning of the eight-week treatment protocol and after offspring euthanasia). The Control group consumed a higher amount of solid diet when compared to the pair-fed and Alcohol groups (50.85 ± 11.25 g) (p < 0.0001). The Pair-fed group ingested all the solid diet offered to them. The food consumption between Pair-fed and Alcohol was equivalent (28.8 ± 9.13 g and 28.02 ± 6.77 g, respectively) (Table 1).
Concerning the liquid diet, although the Pair-fed group received a carbohydrate solution, which is equivalent to an alcohol solution in calories, the rats did not ingest all the solution available to them. The Alcohol group consumed an average of 52.63 ± 15.41 mL of alcohol solution, and the Pair-fed group ingested 34.6 ± 11.95 mL of carbohydrate solution (p < 0.0001). In the liquid diet, the Control group also showed higher consumption (91.67 ± 18.74 mL) than the alcohol and Pair-fed groups (p < 0.0001) (Table 1).
Weight gain was observed in both the control and Alcohol groups (49.89 ± 19.27 % and 29.93 ± 15.21 %, respectively). However, the Pair-fed group showed weight loss (-1.67 % ± 7.46 g) differing significantly from the Alcohol and Control groups (p = 0.0099 and p = 0.0003, respectively).
Determination of blood ethanol concentrations in dams
As previously described [19] the Alcohol group had an average blood ethanol concentration of 1280 ± 730 mg/dL, while the Pair-fed and Control groups showed negative alcohol concentration in the blood.
qRT-PCR
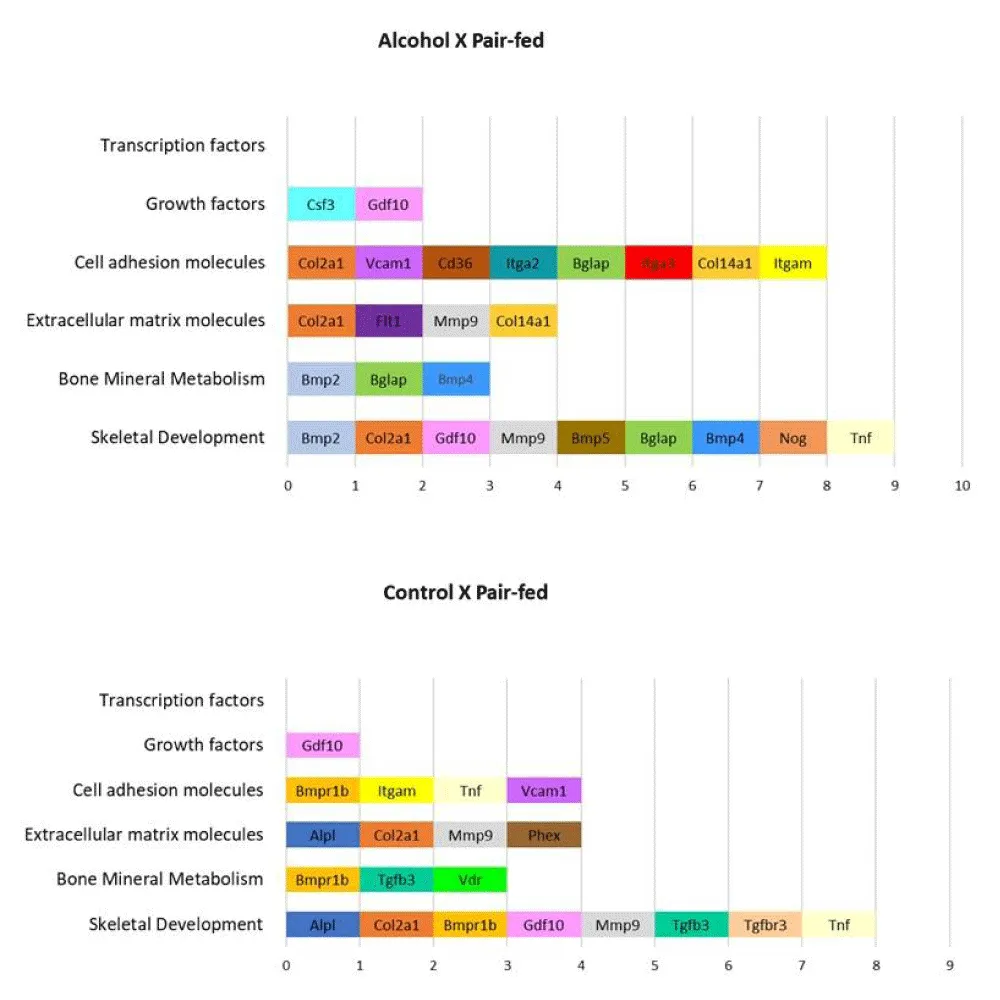
TThe threshold cycle (Ct) values predominantly fell below 25 or were observed between 25 and 30 across all experimental groups, thereby confirming the reliability of this study. Beta-2 microglobulin (B2m) was selected as a normalizing reference. The bone gene set allows the separation of genes by category according to the biological events of osteogenesis: skeletal development, bone mineral metabolism, extracellular matrix molecules (ECM), cell adhesion molecules, growth factors, and transcription factors (Qiagen company catalog provides this separation of genes by category according to the biological events of osteogenesis, Table 2 – Supplementary Material). A first general analysis showed expression variation in 23 bone genes that fall into 3 main categories: Skeletal development (38,2%), Cell adhesion molecules (23,5%), and Bone mineral metabolism (20,5%). Extracellular matrix molecules correspond to 11.7% of the modulated genes. Growth and transcription factors correspond to 2.9 % each. Few genes are related to more than one category. A diagram was provided to categorize the biological events associated with the genes that exhibited differential expression (Figure 2).
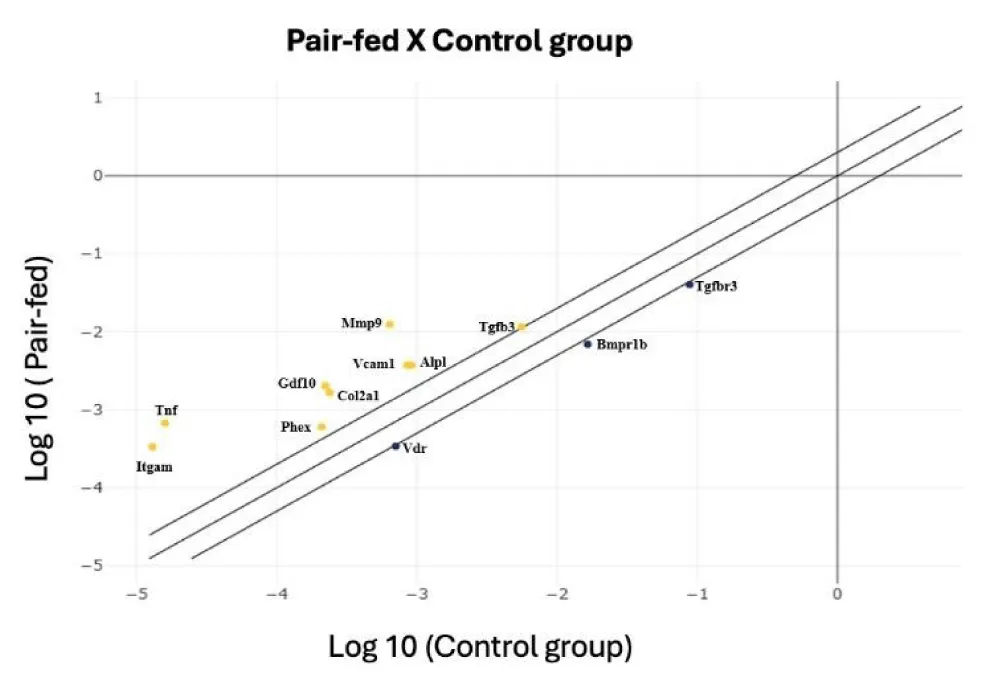
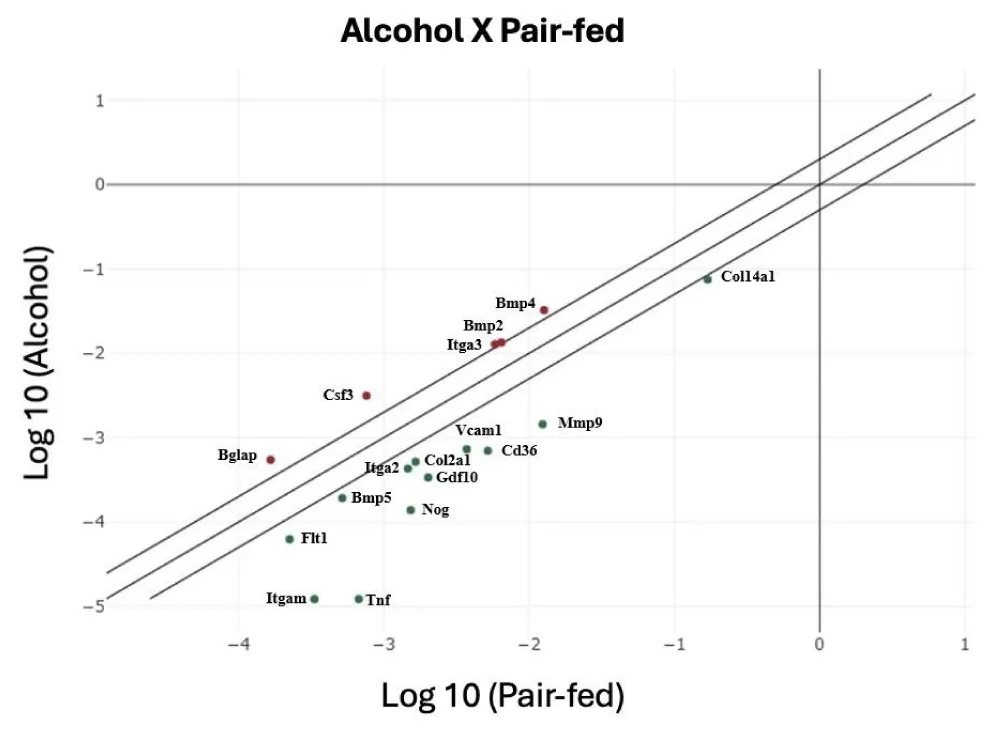
In comparison to the Control group, the Pair-fed group exhibited significant (p < 0.05) downregulation of the Bmpr1b, Tgfbr3, and Vdr genes and upregulation of Alpl, Col2a1, Gdf10, Itgam, Mmp9, Phex, Tgfb3, Tnf, and Vcam1 genes (Table 2 and Figure 3). Out of the 84 analyzed genes associated with skeletal development, growth, and metabolism, 17 genes exhibited significant alterations in their expression when compared between the Alcohol and the Pair-fed groups. Most of these genes demonstrated downregulation in the Alcohol group relative to the Pair-fed group, including Bmp5, Cd36, Col14a1, Col2a1, Flt1, Gdf10, Itga2, Itgam, Mmp9, Nog, Tnf, and Vcam1. Conversely, Bgla, Bmp2, Bmp4, Csf3, and Itga3 were upregulated in the Alcohol group compared to the Pair-fed group. Detailed expression and statistical results for all genes are presented in Table 2 and Figure 4 (the comparative analysis between the Control and Alcohol groups is presented in the supplementary material - Table 3).
Discussions
It is well accepted that increased alcohol consumption has harmful effects on bone growth and metabolism, leading to reduced bone density and altered architecture that elevates the fracture risk in adult ages [42-44]. These effects are either directly or indirectly influenced by various cell types, hormones, and growth factors that govern bone metabolism [45-47]. Exposure to alcohol during prenatal development also contributes to enduring negative health consequences [48]. Studies in animal models showed various disturbances in bone growth and quality due to prenatal alcohol exposure [7-12,14-17,25].
Osteoblasts are one bone cell type that is directly related to bone development, growth, and function. It is well known that osteoblasts and osteogenic stem cells (pre-osteoblasts) treated with ethanol modify their phenotype, showing altered proliferation and differentiation [18,49,50], which may be the result of an altered gene expression pattern, but the molecular mechanisms of alcohol action on osteoblast function are unclear. Here, the observation that genes related to bone development, metabolism, and function have their expression modulated by maternal alcohol ingestion sheds light on the molecular effect of prenatal alcohol exposure on osteoblasts.
A previous study from our group performed with the same animals and consequently with the same conditions as the present work showed that alcohol is genotoxic for the osteoblasts [19]. It is important to emphasize that we have measured the blood alcohol concentration in rat dams, and the average volume of ethanol solution ingested by the Alcohol group corresponds to 1280 ± 730 mg/dL blood ethanol concentration. The maternal blood ethanol concentration is rarely known in human studies and is often not measured in animal studies, making it difficult to compare among studies. The measured values in the present study are considered a high ethanol dosage because even the lower measured values (550 mg/dL blood ethanol concentration) are still considered a high ethanol dosage [14,51-53]. In this way, the prenatal alcohol exposure was able to damage the DNA of osteoblasts, suggesting a direct action of alcohol on these cells and consequently on bone formation [19]. As expected, the same prenatal alcohol exposure was also able to modulate osteoblast gene expression. Taken together, these findings indicate that alcohol changes the bone tissue by affecting the osteoblasts.
Osteoblasts and osteoclasts are the pivotal cells involved in bone turnover: they are responsible for bone formation and bone resorption, respectively. In particular, osteoblasts arise from the osteogenic differentiation of mesenchymal stem cells through a process regulated in different steps. Osteocytes, other important skeletal cells that derive from mature osteoblasts and are surrounded by secreted extracellular matrix, regulate osteoblast and osteoclast activity and, consequently, maintain bone homeostasis. Alterations of mesenchymal stem cells may occur during commitment or differentiation towards the osteogenic lineage, causing demineralization or bone loss in different pathological settings [54].
Within the array of genes participating in osteogenic differentiation, certain ones are already recognized for their specific functions in this intricate process. For this reason, some genes are more evaluated in studies on bone formation, like osteopontin (Spp1), alkaline phosphatase (Alpl), and osteocalcin (Bglap), which are the most investigated osteogenic-related genes [55,56]. In the present study, 84 genes related to osteogenesis were evaluated in osteoblasts differentiated in vitro (primary pre-osteoblasts obtained from the calvaria of fetuses from the three groups). Variations in gene expression were evaluated by real-time PCR and clustered in categories: skeletal development, bone mineral metabolism, extracellular matrix molecules (ECM), cell adhesion molecules, growth factors, and transcription factors. The results showed that alcohol alters the genes expression in practically all biological events related to osteogenesis. Among the 17 genes that showed significant changes in gene expression between the Alcohol group compared to the Pair-fed group, 12 genes presented downregulation. The Bmp5, Col2a1, Gdf10, Mmp9, Nog, and Tnf have been observed in skeletal development. The Cd36, Itga2, Itgam, and Vcam1 genes were found in association with cell adhesion molecules. The Col14a1, Col2a1, and Flt1 genes have been related to extracellular matrix molecules. In contrast, the Bglap, Bmp2, Bmp4, Csf3, and Itga3 genes were upregulated in the Alcohol group compared to the Pair-fed group. Changes in Bglap, Bmp2, and Bmp4 gene expression are related to skeletal development and bone mineral metabolism. The Csf3 is a growth factor, and the Bglap and Itga3 genes are usually involved with cell adhesion, according to the manufacturer’s description of the plate model chosen (Rat Osteogenesis (PARN 026ZA_12/ Figure 2).
Osteocalcin, also referred to as bone gamma-carboxyglutamate (gla) protein, is a non-collagenous protein primarily found in bone tissue and is notably up-regulated during the later stages of cellular differentiation. This stage coincides with the onset of mineralization, suggesting that osteocalcin may play a part in the regulation of matrix mineralization [57]. In a prior investigation by our research team [18], the impact of alcohol was explored through assessments of cell adhesion, proliferation, viability, total protein content, Alkaline Phosphatase activity, and bone matrix formation. The findings indicated that prenatal ethanol exposure enhances cell differentiation, leading to increased alkaline phosphatase production and mineralized bone matrix formation on the 14th and 21st days of cultivation. The current study reveals an overexpression of the osteocalcin gene (Bglap) in the Alcohol group. Taken together, these outcomes suggest that the upregulation of this gene may be associated with heightened matrix mineralization, as observed in our prior work [18]. This overexpression of the Bglap gene likely contributes to accelerated mineralization noted in our previous investigation. Studies have shown that animals exposed to prenatal alcohol exhibit smaller bones or reduced stature [7,8,11-15,23,24,43]. One potential mechanism behind this phenomenon could be the acceleration of mineralization due to alcohol-induced overexpression of Bglap, as demonstrated in this study, resulting in faster bone mineralization and, consequently, premature cessation of bone growth.
Numerous cytokines and growth factors play pivotal roles in bone formation, with BMPs (bone morphogenetic proteins) particularly central to skeletogenesis. BMPs contribute to mesenchyme condensation, skeleton morphogenesis, growth plate development, osteoblast differentiation, and the regulation of postnatal bone and cartilage maintenance [58,59]. Among the 14 known BMPs, BMP-2, 4, 5, 6, 7, and 9 exhibit potent osteogenic activity [58]. Our results demonstrated prenatal alcohol exposure alters the expression of BMPs, with Bmp5 showing reduced expression while Bmp2 and Bmp4 are overexpressed. This suggests that alcohol directly affects osteoblasts, influencing their differentiation process. In addition, BMP-2 is particularly significant as it significantly enhances Bglap expression, with short-term BMP-2 expression being crucial for inducing irreversible bone formation [58]. Considering these findings and the results of Bglap and BMP-2 expression in our study, it can be inferred that prenatal alcohol exposure upregulates these genes, promoting bone formation. This supports the hypothesis of alcohol’s mechanism in bone formation and its role in accelerating bone mineralization, potentially contributing to the shorter stature observed in descendants of alcohol consumers.
The Growth differentiation factor 10 (Gdf10), also referred to as bone morphogenetic protein 3B (Bmp3b), belongs to the transforming growth factor beta (TGF-ß) superfamily [58,60]. Gdf10 is strongly expressed in developing skeletal structures in embryos and in bones, the brain (especially the cerebellum), the aorta, and adipose tissues in adult rodents [61,62]. Moreover, Gdf10 plays a role in inhibiting osteoblast differentiation [63] and is crucial for head formation in Xenopus embryos [64]. In our study, the alcohol-exposed group demonstrated reduced expression of the Gdf10 gene, supporting the hypothesis of alcohol’s direct impact on osteoblasts and bone development. Furthermore, we can infer its action, particularly in skull bone formation, given the essential role of this gene in head development.
In the present study, cells from animals exposed to prenatal alcohol consumption exhibited reduced expression of two genes associated with the extracellular matrix (ECM) structure, Col2a1 and Col14a1. Type II collagen (Col2a1) is a key component of the cartilage matrix. Along with other proteins and proteoglycans, Col2a1 forms complex extracellular scaffolds that support mechanical forces, maintain physiological homeostasis, and provide anchoring sites for chondrocytes, extracellular matrix molecules, and growth factors [65]. Beyond its structural role, Col2a1 serves as an important extracellular signaling molecule, regulating chondrocyte proliferation, metabolism, and differentiation [66]. The degradation and reduction of Col2a1 are considered characteristic pathological markers [67-69]. Type- XIV collagen (Col14a1) is often present in areas of high mechanical stress [70-74].
Previous research 74 revealed that mice lacking Col14a1 are viable. However, detailed examinations of their skin and tendons showed defects in fibril growth and fiber assembly during embryonic development. Due to altered fibrillogenesis, the fiber structure was compromised, leading to significantly reduced biomechanical functions in these tissues [74]. This suggests that Col14a1 is crucial for ECM assembly and tissue function in tendons and skin. This way, our findings suggest that prenatal alcohol exposure also impacts the bone ECM.
In the current study, the expression of six genes associated with cell adhesion—Itga2, Itgam, Itga3, Cd36, Vcam-1, and Bglap—was found to be altered. Cell adhesion is essential for all multicellular organisms, enabling interaction and coordination within cell populations.
Integrins, a key class of cell adhesion receptors, play a vital role in “maintaining the integrity of the cytoskeletal- extracellular matrix linkage.” This concept was first established in the 1970s and 1980s through the research of Erkki Ruoslahti and Richard O. Hynes, who are recognized as the pioneers of integrin research [75]. Integrins consist of an alpha (a) subunit and a beta (b) subunit, each with a specific affinity for different ECM components. The integrin family comprises 24 known members, which play active roles in regulating cellular growth, differentiation, and apoptosis. Each integrin heterodimer performs distinct functions in specific contexts, although some functions partially overlap with those of other integrin family members [76]. Our results revealed that the Itga2 and Itgam genes were underexpressed, while the Itga3 gene was overexpressed in the group exposed to alcohol during pregnancy. This corroborates our findings that prenatal alcohol exposure also affects the bone ECM.
CD36 is a glycoprotein embedded within cell membranes and is found in a variety of immune and non-immune cells, including bone cells [77-79]. It interacts with various external ligands, implicating this receptor in diverse biological processes. While the involvement of Cd36 in various physiological functions is well-documented, its specific role in osteoblasts remains to be fully understood. Kervokova et al. (2013) investigated Cd36’s impact on bone metabolism and osteoblast activities, revealing that mice lacking Cd36 exhibited an osteopenic phenotype in trabecular bone. In vitro studies on bone marrow-derived mesenchymal stem cells and osteoblasts from Cd36-deficient mice showed decreased cell viability and expansion. Additionally, key osteoblastic transcription factors like Runx2 and Osterix, along with osteocalcin and bone sialoprotein expression, were downregulated in Cd36-deficient cells. These findings suggest that Cd36 plays a crucial role in bone metabolism, ensuring proper bone formation. In our current investigation, we observed reduced expression of the Cd36 gene in the Alcohol group. This outcome underscores the impact of prenatal alcohol exposure on osteoblasts, potentially disrupting normal bone formation processes.
Vascular cell adhesion molecule 1 (Vcam-1) was initially recognized as a glycoprotein on the surface of endothelial cells in 1989. This glycoprotein is alternatively referred to as CD106 and can be triggered for expression. Vcam-1 expression is activated by pro-inflammatory cytokines, including Tnfa, and also by ROS, oxidized low-density lipoprotein, high glucose concentration, toll-like receptor agonists, and shear stress. Under high levels of inflammation and chronic conditions in some diseases, Vcam-1 is also expressed on the surface of other cells, including tissue macrophages, dendritic cells, bone marrow fibroblasts, myoblasts, oocytes, Kupffer cells, Sertoli cells, and cancer cells [80]. In this study, the Vcam-1 gene exhibited reduced expression following prenatal alcohol exposure, likely due to the underexpression of the Tnf gene, which activates this protein.
Tumor necrosis factor alpha (Tnfa) functions as a pro-inflammatory cytokine, prompting the expression of various inflammatory molecules, including other cytokines and cell adhesion molecules [80]. While Tnf’s impact on bone tissue, particularly in bone resorption, is established [81], its effect on the differentiation of mesenchymal stromal cells into osteoblasts remains contentious. Some research indicates Tnf inhibits this differentiation, while others suggest it can stimulate osteogenic differentiation by increasing levels of Runx2, Osterix, osteocalcin, Bmp-2, and alkaline phosphatase [81]. The conflicting roles of Tnf in mesenchymal stromal cell osteogenic differentiation likely stem from differences in cellular stages, Tnf concentration, and exposure duration. In our current investigation, we noted a decrease in the expression of the Tnf gene and an increase in the expression of the osteocalcin gene. This observation further supports the previously observed trend of positive regulation identified in other studies.
Flt1, also known as Fms Related Receptor Tyrosine Kinase 1, is a protein-coding gene that is alternatively referred to as Vegfr1 (Vascular Endothelial Growth Factor Receptor 1). It encodes a member of the vascular endothelial growth factor receptor (VEGFR) family. Expression of this protein is observed across various non-endothelial cell types, including vascular smooth muscle cells and macrophages [82]. While the primary focus of research on Flt1 has centered around angiogenesis, studies have also noted its expression in other cell types, such as osteoblasts and osteoclasts [83-86]. Accumulating evidence suggests potential implications of the VEGF/Flt-1 system in bone formation [87,88]. Recently, Xu and colleagues [89] demonstrated that Sod3 and its downstream gene, Flt1, influence osteogenic and adipogenic differentiation through the PI3K/AKT and MAPK pathways, ultimately impacting bone mass. In the current study, the alcohol-exposed group exhibited reduced expression of the Flt1 gene, underscoring the effect of prenatal alcohol exposure on osteogenic differentiation.
The Csf3 gene encodes granulocyte colony-stimulating factor (GCSF), a cytokine that functions as a hematopoietic growth factor governing the survival, growth, and specialization of granulocyte precursors and neutrophil activity [90]. Despite lacking a direct physiological role in bone regulation, GCSF exhibits pharmacological impact on the skeleton in vivo [91]. The mechanisms by which pharmacological GCSF acts on the osteoblast lineage or osteoclasts have not been fully resolved; however, it is clear that the pharmacological effects of GCSF on the skeleton result from indirect action on the osteoblast lineage, as the GCSF receptor is not expressed by osteoblasts or osteocytes [92,93]. In the current study, the cells exposed to alcohol during development displayed heightened expression of the Csf3 gene; this finding indicates that alcohol exerts an indirect influence on osteoblasts as well.
Matrix metalloproteinases (MMPs) are zinc-dependent enzymes that play a crucial role in the regeneration of the ECM by degrading its components. These proteins target and cleave structural elements of the ECM, such as collagen and gelatin, facilitating its breakdown and subsequent regeneration [94]. Under both physiological and pathological conditions, bone cells actively express MMPs, which are believed to be vital for the viability and functionality of osteoclasts, osteoblasts, and osteocytes. Additionally, these MMPs are crucial for the formation and development of chondrocytes, all of which are influenced by bone ECM [94]. Given the dynamic nature of bone tissue and the necessity for various enzymes to degrade the organic components of the bone matrix, the actions
In the current study, the expression of six genes associated with cell adhesion—Itga2, Itgam, Itga3, Cd36, Vcam-1 and Bglap—was found to be altered. Cell adhesion is essential for all multicellular organisms, enabling interaction and coordination within cell populations.
Integrins, a key class of cell adhesion receptors, play a vital role in “maintaining the integrity of the cytoskeletal- extracellular matrix linkage.” This concept was first established in the 1970s and 1980s through the research of Erkki Ruoslahti and Richard O. Hynes, who are recognized as the pioneers of integrin research [75]. Integrins consist of an alpha (a) subunit and a beta (b) subunit, each with a specific affinity for different ECM components. The integrin family comprises 24 known members, which play active roles in regulating cellular growth, differentiation, and apoptosis. Each integrin heterodimer performs distinct functions in specific contexts, although some functions partially overlap with those of other integrin family members [76]. Our results revealed that the Itga2 and Itgam genes were underexpressed, while the Itga3 gene was overexpressed in the group exposed to alcohol during pregnancy. This corroborates our findings that prenatal alcohol exposure also affects the bone ECM.
CD36 is a glycoprotein embedded within cell membranes and is found in a variety of immune and non-immune cells, including bone cells [77-79]. It interacts with various external ligands, implicating this receptor in diverse biological processes. While the involvement of Cd36 in various physiological functions is well-documented, its specific role in osteoblasts remains to be fully understood. Kervokova et al. (2013) investigated Cd36’s impact on bone metabolism and osteoblast activities, revealing that mice lacking Cd36 exhibited an osteopenic phenotype in trabecular bone. In vitro studies on bone marrow-derived mesenchymal stem cells and osteoblasts from Cd36-deficient mice showed decreased cell viability and expansion. Additionally, key osteoblastic transcription factors like Runx2 and Osterix, along with osteocalcin and bone sialoprotein expression, were downregulated in Cd36-deficient cells. These findings suggest that Cd36 plays a crucial role in bone metabolism, ensuring proper bone formation. In our current investigation, we observed reduced expression of the Cd36 gene in the Alcohol group. This outcome underscores the impact of prenatal alcohol exposure on osteoblasts, potentially disrupting normal bone formation processes.
Vascular cell adhesion molecule 1 (Vcam-1) was initially recognized as a glycoprotein on the surface of endothelial cells in 1989. This glycoprotein is alternatively referred to as CD106 and can be triggered for expression. Vcam-1 expression is activated by pro-inflammatory cytokines, including Tnfa, and also by ROS, oxidized lowdensity lipoprotein, high glucose concentration, toll-like receptor agonists, and shear stress. Under high levels of inflammation and chronic conditions in some diseases, Vcam-1 is also expressed on the surface of other cells, including tissue macrophages, dendritic cells, bone marrow fibroblasts, myoblasts, oocytes, Kupffer cells, Sertoli cells, and cancer cells [80]. In this study, the Vcam-1 gene exhibited reduced expression following prenatal alcohol exposure, likely due to the under expression of the Tnf gene, which activates this protein.
Tumor necrosis factor alpha (Tnfa) functions as a pro-inflammatory cytokine, prompting the expression of various inflammatory molecules, including other cytokines and cell adhesion molecules [80]. While Tnf’s impact on bone tissue, particularly in bone resorption, is established [81], its effect on the differentiation of mesenchymal stromal cells into osteoblasts remains contentious. Some research indicates Tnf inhibits this differentiation, while others suggest it can stimulate osteogenic differentiation by increasing levels of Runx2, Osterix, osteocalcin, Bmp-2, and alkaline phosphatase [81]. The conflicting roles of Tnf in mesenchymal stromal cell osteogenic differentiation likely stem from differences in cellular stages, Tnf concentration, and exposure duration. In our current investigation, we noted a decrease in the expression of Tnf gene and an increase in the expression of the osteocalcin gene. This observation further supports the previously observed trend of positive regulation identified in other studies.
Flt1, also known as Fms Related Receptor Tyrosine Kinase 1, is a protein-coding gene that is alternatively referred to as Vegfr1 (Vascular Endothelial Growth Factor Receptor 1). It encodes a member of the vascular endothelial growth factor receptor (VEGFR) family. Expression of this protein is observed across various non-endothelial cell types, including vascular smooth muscle cells and macrophages [82]. While the primary focus of research on Flt1 has centered around angiogenesis, studies have also noted its expression in other cell types such as osteoblasts and osteoclasts [83-86]. Accumulating evidence suggests potential implications of the VEGF/Flt-1 system in bone formation [87,88]. Recently, Xu and colleagues [89] demonstrated that Sod3 and its downstream gene, Flt1, influence osteogenic and adipogenic differentiation through the PI3K/AKT and MAPK pathways, ultimately impacting bone mass. In the current study, the alcohol-exposed group exhibited reduced expression of the Flt1 gene, underscoring the effect of prenatal alcohol exposure on osteogenic differentiation.
The Csf3 gene encodes granulocyte colony-stimulating factor (GCSF), a cytokine that functions as a hematopoietic growth factor governing the survival, growth, and specialization of granulocyte precursors and neutrophil activity [90]. Despite lacking a direct physiological role in bone regulation, GCSF exhibits pharmacological impact on the skeleton in vivo [91]. The mechanisms by which pharmacological GCSF acts on the osteoblast lineage or osteoclasts have not been fully resolved, however, it is clear that the pharmacological effects of GCSF on the skeleton result from indirect action on the osteoblast lineage, as the GCSF receptor is not expressed by osteoblasts or osteocytes [92,93]. In the current study, the cells exposed to alcohol during development displayed heightened expression of the Csf3 gene; this finding indicates that alcohol exerts an indirect influence on osteoblasts as well.
Matrix metalloproteinases (MMPs) are zinc-dependent enzymes that play a crucial role in the regeneration of the ECM by degrading its components. These proteins target and cleave structural elements of the ECM, such as collagen and gelatin, facilitating its breakdown and subsequent regeneration [94]. Under both physiological and pathological conditions, bone cells actively express MMPs, which are believed to be vital for the viability and functionality of osteoclasts, osteoblasts, and osteocytes. Additionally, these MMPs are crucial for the formation and development of chondrocytes, all of which are influenced by bone ECM [94]. Given the dynamic nature of bone tissue and the necessity for various enzymes to degrade the organic components of the bone matrix, the actions of MMPs and their inhibitors hold significant physiological importance, in this manner, can state that MMPs are crucial mediators of bone physiology [94]. The literature indicates that the expression of matrix metalloproteinase-9 (MMP-9) varies as a child grows, and this enzyme is expressed in trophoblasts and osteoclasts during early development, suggesting its role in bone implantation and resorption [94]. Furthermore, the absence of certain molecules, including MMP-9, during skeletal development lead to significant defects in long bone plates, impeding normal bone production [94]. In the present study, alcohol caused in the under expression of the Mmp-9 gene. This finding provides further compelling evidence that alcohol impacts the ECM, thereby influencing the bone development of offspring whose mothers consumed alcohol during pregnancy.
Noggin, also referred to as Nog and encoded by the Nog gene, is a secreted homodimeric glycoprotein involved in the development of various body tissues, including nerve tissue, muscles, and bones [95]. In bone tissue, Nog regulates bone morphogenetic protein signaling by preventing these proteins from binding to cell receptors, thereby inhibiting signal transduction. By this mechanism, it modulates various signaling pathways during cartilage and bone formation in human development [95]. This gene is considered very important for bone development, as mutations in the Nog gene result in alterations in bone morphogenetic protein signaling, leading to different types of dysplasias [96-99]. Furthermore, some studies suggest Nog as a potential candidate for craniofacial disorders [100-102]. Our results showed that prenatal alcohol exposure led to reduced expression of the Nog gene, which may have influenced the expression of the Bmp2, Bmp4, and Bmp5 genes, as changes in their expression were observed in the alcohol-exposed group. Additionally, literature suggests that the Nog gene plays a role in head formation; thus, the under expression of this gene in our study indicates that the cranial bone changes observed in previous studies may be related to this gene alteration [16,21,22,24].
It is well known that alcohol has low nutritional value, however its metabolism by the liver provides energy to the body (7.1 kcal/g), leading to a sense of satiety. Thus, nutritionally important foods are replaced by alcohol, which commonly leads to malnutrition due to the lower intake of nutrients [103,104]. In agreement, in our study, there was a significant decrease in solid food intake by the animals of the Alcohol group when compared to the Control group. Considering that the Alcohol group ingested fewer nutrients, the presence of the Pair-fed group was important to control this variable. Regarding nutrient deficiency, mimicked by the Pair-fed group, our results showed alterations in the gene expression of 12 genes evaluated. The Bmpr1b, Tgfbr3 and Vdr genes were downregulated in Pair-fed group compared to Control group. These genes have been observed in cell adhesion molecules (Bmpr1b), skeletal development (Bmpr1b and Tgfbr3) and bone mineral metabolism (Bmpr1b and Vdr). On the other hand, the Alpl, Col2a1, Gdf10, Itgam, Mmp9, Phex, Tgfb3, Tnf and Vcam1 genes were upregulated in the Pair-fed group compared to the Control group. The Alpl, Col2a1, Gdf10, Mmp9, Tgfb3 and Tnf genes are important for skeletal development, while the Itgam and Vcam1 genes have been associated with cell adhesion and Phex gene have been associated bone mineral metabolism (Figure 2).
In general, our findings suggest that this direct action on osteoblast gene expression may be part of the mechanism involved in arrested development and impaired skeletal function due to prenatal alcohol exposure. The mechanisms underlying alcohol’s impact on bone formation remain unclear. Examining gene expression linked to osteogenic differentiation holds the potential to unravel these mechanisms in bone development, given that osteoblasts and ECM are two pivotal elements in the process [105].
Although the methodology adopted for the treatment of the animals in our study is well described and previously performed by other authors [14,16,18,19,27,51], some limitations should be pointed out.
Firstly, the liquid diet of isocaloric group (Pair-fed group) was not strictly controlled, although nutritional pairing was accomplished regarding the solid food. Moreover, the extent of the pups’ exposure to alcohol during the breastfeeding period was not assessed. Despite these limitations, this kind of methodology have been well accepted in the scientific environment [14,16,18,19,26,27,51].
Another limitation that should be highlighted is the absence of individual sample evaluation for each animal. Since we have evaluated the cells jointly and not individually, we could not check the individual gene expression or difference between males and females. However, our findings represent a screening for selection of some genes that should be investigated individually in male and female newborns using the same model where the cells are extracted from the calvariae of newborns while still in the pre-osteoblastic mesenchymal cell stage. In our model, the cells are allowed to differentiate into osteoblasts during in vitro cultivation, without the addition of alcohol to the culture medium. This approach proved to be suitable to assess the influence of alcohol on gene expression during fetal development, demonstrating a direct effect of alcohol on bone development. Future in vitro and in vivo studies should also explore osteoclast behavior to assess the full effect of maternal ingestion on bone turnover of the newborns.
In summary, our study evaluated the gene expression of osteoblasts from the calvaria of newborn rats whose mothers received alcohol prior to breeding and during the three weeks of gestation. A suitable alcohol level could be detected in the blood of these mothers. As expected, the animals from the Alcohol group exhibited a tendency to weight gain, even with lower ingestion of food. The poor nutritional intake of the Alcohol group was compensated with the insertion of the Pair-feed group which received lower nutrients. After the treatment of the mothers, pre-osteoblast of the newborns was extracted from the calvaria and differentiated into primary osteoblasts that were posteriorly evaluated regarding osteogenic genes. Altered expressions were detected in 17 genes of which some were downregulated and others upregulated in the Alcohol group.
Conclusion
This study demonstrate that ethanol directly affects fetal osteoblasts by altering gene expression related to osteogenic differentiation in newborn rats, including Itgam, Nog, Tnf, and Mmp-9. Such alterations may impair overall fetal skeletal development. This study highlights the potential repercussions of maternal alcohol consumption on the skeletal health of offspring, paving the way for further investigations into its long-term effects.
Acknowledgments
With appreciation for the São José dos Campos Institute of Science and Technology, UNESP – São Paulo State University animal care and use laboratory.
Funding
This research work was supported by FAPESP (São Paulo Research Foundation - Fundação de Amparo à Pesquisa do Estado de São Paulo - Grant number: 2012/10643-3). Isabel C. S. Carvalho received a scholarship from the Brazilian governmental research agency, CAPES (Coordination for the Improvement of Higher Education Personnel - Coordenação de Aperfeiçoamento de Pessoal de Nível Superior).
Conflict of interest statement
The authors report no conflicts of interest related to this study.
Data availability statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Ethics approval statement
This study was approved by the ethics committee of São José dos Campos Institute of Science and Technology, UNESP – São Paulo State University (Protocol No. 01/2012-PA/CEP).
- Barr HM, Streissguth AP. Identifying maternal self-reported alcohol use associated with fetal alcohol spectrum disorders. Alcohol Clin Exp Res. 2001;25(2):283-287. Available from: https://psycnet.apa.org/doi/10.1111/j.1530-0277.2001.tb02210.x
- Hoyme HE, May PA, Kalberg WO, Kodituwakku P, Gossage JP, Trujillo PM, et al. A practical clinical approach to diagnosis of fetal alcohol spectrum disorders: clarification of the 1996 Institute of Medicine criteria. Pediatrics. 2005;115(1):39-47. Available from: https://doi.org/10.1542/peds.2004-0259
- O’Leary CM, Elliott EJ, Nassar N, Bower C. Exploring the potential to use data linkage for investigating the relationship between birth defects and prenatal alcohol exposure. Birth Defects Res A Clin Mol Teratol. 2013;97(7):497-504. Available from: https://doi.org/10.1002/bdra.23142
- Hoyme HE, Kalberg WO, Elliott AJ, Blankenship J, Buckley D, Marais A, et al. Updated clinical guidelines for diagnosing fetal alcohol spectrum disorders. Pediatrics. 2016;138(2):e20154256. Available from: https://doi.org/10.1542/peds.2015-4256
- Chung DD, Pinson MR, Bhenderu LS, Lai MS, Patel RA, Miranda RC. Toxic and teratogenic effects of prenatal alcohol exposure on fetal development, adolescence, and adulthood. Int J Mol Sci. 2021;22(16):8785. Available from: https://doi.org/10.3390/ijms22168785
- Charness ME. Fetal alcohol spectrum disorders: awareness to insight in just 50 years. Alcohol Res. 2022;42(1):05. Available from: https://doi.org/10.35946/arcr.v42.1.05
- Day NL, Richardson G, Robles N, Sambamoorthi U, Taylor P, Scher M, et al. Effect of prenatal alcohol exposure on growth and morphology of offspring at 8 months of age. Pediatrics. 1990;85(5):748-752. Available from: https://doi.org/10.1542/peds.85.5.748
- Day NL, Leech SL, Richardson GA, Cornelius MD, Robles N, Larkby C. Prenatal alcohol exposure predicts continued deficits in offspring size at 14 years of age. Alcohol Clin Exp Res. 2002;26(10):1584-1591. Available from: https://doi.org/10.1097/01.alc.0000034036.75248.d9
- Keiver K, Herbert L, Weinberg J. Effect of maternal ethanol consumption on maternal and fetal calcium metabolism. Alcohol Clin Exp Res. 1996;20(7):1305-1312. Available from: https://doi.org/10.1111/j.1530-0277.1996.tb01127.x
- Keiver K, Weinberg J. Effect of duration of maternal alcohol consumption on calcium metabolism and bone in the fetal rat. Alcohol Clin Exp Res. 2004;28(3):456-467. Available from: https://doi.org/10.1097/01.alc.0000118312.38204.c5
- Lee M, Leichter J. Effect of litter size on the physical growth and maturation of the offspring of rats given alcohol during gestation. Growth. 1980;44(4):327-335. Available from: https://scienceon.kisti.re.kr/srch/selectPORSrchArticle.do?cn=NART34782820
- Lee M, Leichter J. Skeletal development in fetuses of rats consuming alcohol during gestation. Growth. 1983;47(3):254-262. Available from: https://pubmed.ncbi.nlm.nih.gov/6642247/
- Ramadoss J, Hogan HA, Given JC, West JR, Cudd TA. Binge alcohol exposure during all three trimesters alters bone strength and growth in fetal sheep. Alcohol. 2006;38(3):185-192. Available from: https://doi.org/10.1016/j.alcohol.2006.06.004
- Simpson ME, Duggal S, Keiver K. Prenatal ethanol exposure has differential effects on fetal growth and skeletal ossification. Bone. 2005;36(3):521-532. Available from: https://doi.org/10.1016/j.bone.2004.11.011
- Weinberg J, D’Alquen G, Bezio S. Interactive effects of ethanol intake and maternal nutritional status on skeletal development of fetal rats. Alcohol. 1990;7(5):383-388. Available from: https://doi.org/10.1016/0741-8329(90)90020-d
- Carvalho ICS, Martinelli C da SM, Milhan NVM, da S Marchini AMP, Dutra TP, de Souza DM, et al. Prenatal alcohol exposure reduces mandibular calcium and phosphorus concentrations in newborn rats. J Oral Sci. 2016;58(3):439-444. Available from: https://doi.org/10.2334/josnusd.16-0061
- Keiver K, Ellis L, Anzarut A, Weinberg J. Effect of prenatal ethanol exposure on fetal calcium metabolism. Alcohol Clin Exp Res. 1997;21(9):1612-1618. Available from: https://doi.org/10.1111/j.1530-0277.1997.tb04497.x
- Carvalho ICS, De Andrade DP, Milhan NVM, de Souza Santos EL, Soares CP, da Rocha RF, et al. Prenatal ethanol exposure affects the proliferation and differentiation of the osteoblasts from newborn rats. Online J Biol Sci. 2015;15(3):134-142. Available from: https://doi.org/10.3844/ojbsci.2015.134.142
- Carvalho ICS, Dutra TP, De Andrade DP, Balducci I, Pacheco-Soares C, da Rocha RF. High doses of alcohol during pregnancy cause DNA damages in osteoblasts of newborn rats. Birth Defects Res A Clin Mol Teratol. 2016;106(2):122-132. Available from: https://doi.org/10.1002/bdra.23468
- Keiver K, Herbert L, Weinberg J. Effect of maternal ethanol consumption on maternal and fetal calcium metabolism. Alcohol Clin Exp Res. 1996;20(7):1305-1312. Available from: https://doi.org/10.1111/j.1530-0277.1996.tb01127.x
- Oyedele OO, Kramer B. Acute ethanol administration causes malformations but does not affect cranial morphometry in neonatal mice. Alcohol. 2008;42(1):21-27. Available from: https://doi.org/10.1016/j.alcohol.2007.10.007
- Anthony B, Vinci-Booher S, Wetherill L, Ward R, Goodlett C, Zhou FC. Alcohol-induced facial dysmorphology in C57BL/6 mouse models of fetal alcohol spectrum disorder. Alcohol. 2010;44(7-8):659-671. Available from: https://doi.org/10.1016/j.alcohol.2010.04.002
- Probyn ME, Zanini S, Ward LC, Bertram JF, Moritz KM. A rodent model of low- to moderate-dose ethanol consumption during pregnancy: patterns of ethanol consumption and effects on fetal and offspring growth. Reprod Fertil Dev. 2012;24(6):859-870. Available from: https://doi.org/10.1071/rd11200
- Shen L, Ai H, Liang Y, Ren X, Anthony CB, Goodlett CR, et al. Effect of prenatal alcohol exposure on bony craniofacial development: a mouse MicroCT study. Alcohol. 2013;47(5):405-415. Available from: https://doi.org/10.1016/j.alcohol.2013.04.005
- Snow ME, Keiver K. Prenatal ethanol exposure disrupts the histological stages of fetal bone development. Bone. 2007;41(2):181-187. Available from: https://doi.org/10.1016/j.bone.2007.04.182
- Marchini AMPS, Deco CP, Lodi KB, Marchini L, Santo AME, Rocha RF. Influence of chronic alcoholism and oestrogen deficiency on the variation of stoichiometry of hydroxyapatite within alveolar bone crest of rats. Arch Oral Biol. 2012;57(10):1385-1394. Available from: https://doi.org/10.1016/j.archoralbio.2012.04.011
- Marchini AMPDS, Gonçalves LL, Salgado MCM, do Prado RF, Marchini L, Carvalho YR, et al. Alcoholic and isocaloric diet, but not ovariectomy, influence the apoptosis of bone cells within the alveolar bone crest of rats. Arch Oral Biol. 2014;59(4):424-433. Available from: https://doi.org/10.1016/j.archoralbio.2014.01.008
- Wiener SG, Shoemaker WJ, Koda LY, Bloom FE. Interaction of ethanol and nutrition during gestation: influence on maternal and offspring development in the rat. J Pharmacol Exp Ther. 1981;216(3):572-579. Available from: https://doi.org/10.1016/S0022-3565(25)32467-5
- Geißler U, Hempel U, Wolf C, Scharnweber D, Worch H, Wenzel KW. Collagen type I-coating of Ti6Al4V promotes adhesion of osteoblasts. J Biomed Mater Res. 2000;51(4):752-760. Available from: https://doi.org/10.1002/1097-4636(20000915)51:4%3C752::aid-jbm25%3E3.0.co;2-7
- Nanci A, Zalzal S, Gotoh Y, McKee MD. Ultrastructural characterization and immunolocalization of osteopontin in rat calvarial osteoblast primary cultures. Microsc Res Tech. 1996;33:214-231. Available from: https://doi.org/10.1002/(sici)1097-0029(19960201)33:2%3C214::aid-jemt11%3E3.0.co;2-x
- Lynch MP, Stein JL, Stein GS, Lian JB. The influence of type I collagen on the development and maintenance of the osteoblast phenotype in primary and passaged rat calvarial osteoblasts: modification of expression of genes supporting cell growth, adhesion, and extracellular matrix mineralization. Exp Cell Res. 1995;216(1):35-45. Available from: https://doi.org/10.1006/excr.1995.1005
- Ziolkowska A, Rucinski M, Pucher A, Tortorella C, Nussdorfer GG, Malendowicz LK. Expression of osteoblast marker genes in rat calvarial osteoblast-like cells, and effects of the endocrine disrupters diphenylolpropane, benzophenone-3, resveratrol and silymarin. Chem Biol Interact. 2006;164(3):147-156. Available from: https://doi.org/10.1016/j.cbi.2006.09.009
- Kim B, Lee JH, Jin WJ, Kim HH, Ha H, Lee ZH. Trapidil induces osteogenesis by upregulating the signaling of bone morphogenetic proteins. Cell Signal. 2018;49:68-78. Available from: https://doi.org/10.1016/j.cellsig.2018.06.001
- Kushwaha P, Tripathi AK, Gupta S, Kothari P, Upadhyay A, Ahmad N, et al. Synthesis and study of benzofuran-pyran analogs as BMP-2 targeted osteogenic agents. Eur J Med Chem. 2018;156:103-117. Available from: https://doi.org/10.1016/j.ejmech.2018.06.062
- Kirkwood KL, Dziak R, Bradford PG. Inositol trisphosphate receptor gene expression and hormonal regulation in osteoblast-like cell lines and primary osteoblastic cell cultures. J Bone Miner Res. 1996;11(12):1889-1896. Available from: https://doi.org/10.1002/jbmr.5650111209
- Laiuppa JA, Santillán GE. Effect of combined action of extracellular ATP and elevated calcium on osteogenic differentiation of primary cultures from rat calvaria. J Cell Biochem. 2016;117(11):2658-2668. Available from: https://doi.org/10.1002/jcb.25565
- Laiuppa JA, Santillán GE. Involvement of GSK3/β-catenin in the action of extracellular ATP on differentiation of primary cultures from rat calvaria into osteoblasts. J Cell Biochem. 2018;119(10):8378-8388. Available from: https://doi.org/10.1002/jcb.27037
- González O, Fong KD, Trindade MCD, Warren SM, Longaker MT, Smith RL. Fluid shear stress magnitude, duration, and total applied load regulate gene expression and nitric oxide production in primary calvarial osteoblast cultures. Plast Reconstr Surg. 2008;122(2):419-428. Available from: https://doi.org/10.1097/prs.0b013e31817d5ff1
- Kunze R, Rösler M, Möller S, Schnabelrauch M, Riemer T, Hempel U, et al. Sulfated hyaluronan derivatives reduce the proliferation rate of primary rat calvarial osteoblasts. Glycoconj J. 2010;27(1):151-158. Available from: https://doi.org/10.1007/s10719-009-9270-9
- Cepeda SB, Sandoval MJ, Crescitelli MC, Rauschemberger MB, Massheimer VL. The isoflavone genistein enhances osteoblastogenesis: signaling pathways involved. J Physiol Biochem. 2020;76(1):99-110. Available from: https://doi.org/10.1007/s13105-019-00722-3
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25(4):402-408. Available from: https://doi.org/10.1006/meth.2001.1262
- Sampson HW. Alcohol’s harmful effects on bone. Alcohol Health Res World. 1998;22(3):190-194. Available from: https://pubmed.ncbi.nlm.nih.gov/15706795/
- Rosa RC, Rodrigues WF, Miguel CB, Cardoso FAG, Espindula AP, Oliveira CJF, et al. Chronic consumption of alcohol adversely affects the bone of young rats. Acta Ortop Bras. 2019;27(6):321-324. Available from: https://doi.org/10.1590/1413-785220192706222834
- Godos J, Giampieri F, Chisari E, Micek A, Paladino N, Forbes-Hernández TY, et al. Alcohol consumption, bone mineral density, and risk of osteoporotic fractures: a dose-response meta-analysis. Int J Environ Res Public Health. 2022;19(3):1515. Available from: https://doi.org/10.3390/ijerph19031515
- Laitinen K, Välimäki M. Alcohol and bone. Calcif Tissue Int. 1991;49:S70-S73. Available from: https://doi.org/10.1007/bf02555094
- Guo M, Huang YL, Wu Q, Chai L, Jiang Z, Zeng Y, et al. Chronic ethanol consumption induces osteopenia via activation of osteoblast necroptosis. Oxid Med Cell Longev. 2021;2021. Available from: https://doi.org/10.1155/2021/3027954
- Asada T, Iwata M, Matsuzaki S, Hamakawa H, Sengan S, Noguchi T, et al. Hypercalcemia and hyperphosphatemia associated with 25-OH vitamin D deficiency in an alcoholic patient with normal renal function. J Clin Transl Endocrinol Case Rep. 2022;23:100107. Available from: https://doi.org/10.1016/j.jecr.2022.100107
- Young SL, Gallo LA, Brookes DSK, Hayes N, Maloney M, Liddle K, et al. Altered bone and body composition in children and adolescents with confirmed prenatal alcohol exposure. Bone. 2022;164:116510. Available from: https://doi.org/10.1016/j.bone.2022.116510
- Chavassieux P, Serre CM, Vergnaud P, Delmas PD, Meunier PJ. In vitro evaluation of dose-effects of ethanol on human osteoblastic cells. Bone Miner. 1993;22(2):95-103. Available from: https://doi.org/10.1016/s0169-6009(08)80221-8
- Rosa ML, Beloti MM, Prando N, Queiroz RHC, De Oliveira PT, Rosa AL. Chronic ethanol intake inhibits in vitro osteogenesis induced by osteoblasts differentiated from stem cells. J Appl Toxicol. 2008;28(2):205-211. Available from: https://doi.org/10.1002/jat.1271
- Keiver K, Weinberg J. Effect of duration of alcohol consumption on calcium and bone metabolism during pregnancy in the rat. Alcohol Clin Exp Res. 2003;27(9):1507-1519. Available from: https://doi.org/10.1097/01.alc.0000086063.71754.c1
- Keiver K, Weinberg J. Effect of duration of maternal alcohol consumption on calcium metabolism and bone in the fetal rat. Alcohol Clin Exp Res. 2004;28(3):456-467. Available from: https://doi.org/10.1097/01.alc.0000118312.38204.c5
- Wang R, Martin CD, Lei AL, Hausknecht KA, Ishiwari K, Richards JB, et al. Prenatal ethanol exposure leads to attention deficits in both male and female rats. Front Neurosci. 2020;14. Available from: https://doi.org/10.3389/fnins.2020.00012
- Valenti MT, Carbonare LD, Mottes M. Osteogenic differentiation in healthy and pathological conditions. Int J Mol Sci. 2016;18(1):41. Available from: https://doi.org/10.3390/ijms18010041
- Chen D, Li Z, Li Z, Sun Y, Liu Q, Yang J, et al. Transcriptome analysis of human peri-implant soft tissue and periodontal gingiva: a paired design study. Clin Oral Investig. 2023;27(7):3937-3948. Available from: https://doi.org/10.1007/s00784-023-05017-y
- Bernardi S, Re F, Bosio K, Dey K, Almici C, Malagola M, et al. Chitosan-hydrogel polymeric scaffold acts as an independent primary inducer of osteogenic differentiation in human mesenchymal stromal cells. Materials (Basel). 2020;13(16). Available from: https://doi.org/10.3390/ma13163546
- Caetano-Lopes J, Canhão H, Fonseca JE. Osteoblasts and bone formation. Acta Reumatol Port. 2007;32(2):103-110. Available from: https://doi.org/10.1016/S1569-2590(08)60130-5
- Wu M, Chen G, Li YP. TGF-β and BMP signaling in osteoblast, skeletal development, and bone formation, homeostasis and disease. Bone Res. 2016;4:16009. Available from: https://doi.org/10.1038/boneres.2016.9
- Katagiri T, Watabe T. Bone morphogenetic proteins. Cold Spring Harb Perspect Biol. 2016;8(6):a021899. Available from: https://doi.org/10.1101/cshperspect.a021899
- Yu-Lee LY, Yu G, Lee YC, Lin S, Pan J, Pan T, et al. Osteoblast-secreted factors mediate dormancy of metastatic prostate cancer in the bone via activation of the TGFβRIII-p38MAPK-pS249/T252RB pathway. Cancer Res. 2018;78(11):2911-2924. Available from: https://doi.org/10.1158/0008-5472.can-17-1051
- Zhao R, Lawler AM, Lee SJ. Characterization of GDF-10 expression patterns and null mice. Dev Biol. 1999;212(1):68-79. Available from: https://doi.org/10.1006/dbio.1999.9326
- Hino J, Miyazawa T, Miyazato M, Kangawa K. Bone morphogenetic protein-3b (BMP-3b) is expressed in adipocytes and inhibits adipogenesis as a unique complex. Int J Obes (Lond). 2012;36(5):725-734. Available from: https://doi.org/10.1038/ijo.2011.124
- Matsumoto Y, Otsuka F, Hino J, Miyoshi T, Takano M, Miyazato M, et al. Bone morphogenetic protein-3b (BMP-3b) inhibits osteoblast differentiation via Smad2/3 pathway by counteracting Smad1/5/8 signaling. Mol Cell Endocrinol. 2012;350(1):78-86. Available from: https://doi.org/10.1016/j.mce.2011.11.023
- Hino J, Nishimatsu SI, Nagai T, Matsuo H, Kangawa K, Nohno T. Coordination of BMP-3b and cerberus is required for head formation of Xenopus embryos. Dev Biol. 2003;260(1):138-157. Available from: https://doi.org/10.1016/s0012-1606(03)00223-9
- Aigner T, Stöve J. Collagens - major component of the physiological cartilage matrix, major target of cartilage degeneration, major tool in cartilage repair. Adv Drug Deliv Rev. 2003;55(12):1569-1593. Available from: https://doi.org/10.1016/j.addr.2003.08.009
- Xin W, Heilig J, Paulsson M, Zaucke F. Collagen II regulates chondrocyte integrin expression profile and differentiation. Connect Tissue Res. 2015;56(4):307-314. Available from: https://doi.org/10.3109/03008207.2015.1026965
- Deng H, Huang X, Yuan L. Molecular genetics of the COL2A1-related disorders. Mutat Res Rev Mutat Res. 2016;768:1-13. Available from: https://doi.org/10.1016/j.mrrev.2016.02.003
- Lian C, Wang X, Qiu X, Wu Z, Gao B, Liu L, et al. Collagen type II suppresses articular chondrocyte hypertrophy and osteoarthritis progression by promoting integrin β1-SMAD1 interaction. Bone Res. 2019;7(1):8. Available from: https://doi.org/10.1038/s41413-019-0046-y
- Demal TJ, Scholz T, Schüler H, Olfe J, Fröhlich A, Speth F, et al. Expanding the clinical spectrum of COL2A1 related disorders by a mass-like phenotype. Sci Rep. 2022;12(1):4489. Available from: https://doi.org/10.1038/s41598-022-08476-7
- Wälchli C, Koch M, Chiquet M, Odermatt BF, Trueb B. Tissue-specific expression of the fibril-associated collagens XII and XIV. J Cell Sci. 1994;107(2):669-681. Available from: https://doi.org/10.1242/jcs.107.2.669
- Berthod F, Germain L, Guignard R, Lethias C, Garrone R, Damour O, et al. Differential expression of collagens XII and XIV in human skin and in reconstructed skin. J Invest Dermatol. 1997;108(5):737-742. Available from: https://doi.org/10.1111/1523-1747.ep12292122
- Niyibizi C, Visconti CS, Kavalkovich K, Woo SLY. Collagens in an adult bovine medial collateral ligament: immunofluorescence localization by confocal microscopy reveals that type XIV collagen predominates at the ligament-bone junction. Matrix Biol. 1995;14(9):743-751. Available from: https://doi.org/10.1016/s0945-053x(05)80017-4
- Young BB, Gordon MK, Birk DE. Expression of type XIV collagen in developing chicken tendons: association with assembly and growth of collagen fibrils. Dev Dyn. 2000;217(4):430-439. Available from: https://doi.org/10.1002/(sici)1097-0177(200004)217:4%3C430::aid-dvdy10%3E3.0.co;2-5
- Ansorge HL, Meng X, Zhang G, Veit G, Sun M, Klement JF, et al. Type XIV collagen regulates fibrillogenesis: premature collagen fibril growth and tissue dysfunction in null mice. J Biol Chem. 2009;284(13):8427-8438. Available from: https://doi.org/10.1074/jbc.m805582200
- Hynes RO. The emergence of integrins: a personal and historical perspective. Matrix Biol. 2004;23(6):333-340. Available from: https://doi.org/10.1016/j.matbio.2004.08.001
- Adorno-Cruz V, Liu H. Regulation and functions of integrin α2 in cell adhesion and disease. Genes Dis. 2018;6(1):16-24. Available from: https://doi.org/10.1016/j.gendis.2018.12.003
- Silverstein RL, Febbraio M. CD36, a scavenger receptor involved in immunity, metabolism, angiogenesis, and behavior. Sci Signal. 2009;2(72):re3. Available from: https://doi.org/10.1126/scisignal.272re3
- Brodeur MR, Brissette L, Falstrault L, Luangrath V, Moreau R. Scavenger receptor of class B expressed by osteoblastic cells are implicated in the uptake of cholesteryl ester and estradiol from LDL and HDL3. J Bone Miner Res. 2008;23(3):326-337. Available from: https://doi.org/10.1359/jbmr.071022
- Carron JA, Wagstaff SC, Gallagher JA, Bowler WB. A CD36-binding peptide from thrombospondin-1 can stimulate resorption by osteoclasts in vitro. Biochem Biophys Res Commun. 2000;270(3):1124-1127. Available from: https://doi.org/10.1006/bbrc.2000.2574
- Kong DH, Kim YK, Kim MR, Jang JH, Lee S. Emerging roles of vascular cell adhesion molecule-1 (VCAM-1) in immunological disorders and cancer. Int J Mol Sci. 2018;19(4). Available from: https://doi.org/10.3390/ijms19041057
- Zhao B. TNF and bone remodeling. Curr Osteoporos Rep. 2017;15(3):126-134. Available from: https://doi.org/10.1007/s11914-017-0358-z
- Ji S, Xin H, Li Y, Su EJ. FMS-like tyrosine kinase 1 (FLT1) is a key regulator of fetoplacental endothelial cell migration and angiogenesis. Placenta. 2018;70:7-14. Available from: https://doi.org/10.1016/j.placenta.2018.08.004
- Gerber HP, Vu TH, Ryan AM, Kowalski J, Werb Z, Ferrara N. VEGF couples hypertrophic cartilage remodeling, ossification and angiogenesis during endochondral bone formation. Nat Med. 1999;5(6):623-628. Available from: https://doi.org/10.1038/9467
- Niida S, Kaku M, Amano H, Yoshida H, Kataoka H, Nishikawa S, et al. Vascular endothelial growth factor can substitute for macrophage colony-stimulating factor in the support of osteoclastic bone resorption. J Exp Med. 1999;190(2):293-298. Available from: https://doi.org/10.1084/jem.190.2.293
- Deckers MML, Karperien M, Van Der Bent C, Yamashita T, Papapoulos SE, Löwik CWGM. Expression of vascular endothelial growth factors and their receptors during osteoblast differentiation. Endocrinology. 2000;141(5):1667-1674. Available from: https://doi.org/10.1210/endo.141.5.7458
- Casella I, Feccia T, Chelucci C, Samoggia P, Castelli G, Guerriero R, et al. Autocrine-paracrine VEGF loops potentiate the maturation of megakaryocytic precursors through Flt1 receptor. Blood. 2003;101(4):1316-1323. Available from: https://doi.org/10.1182/blood-2002-07-2184
- Uchida S, Sakai A, Kudo H, Otomo H, Watanuki M, Tanaka M, et al. Vascular endothelial growth factor is expressed along with its receptors during the healing process of bone and bone marrow after drill-hole injury in rats. Bone. 2003;32(5):491-501. Available from: https://doi.org/10.1016/s8756-3282(03)00053-x
- Otomo H, Sakai A, Uchida S, Tanaka S, Watanuki M, Moriwaki S, et al. Flt-1 tyrosine kinase-deficient homozygous mice result in decreased trabecular bone volume with reduced osteogenic potential. Bone. 2007;40(6):1494-1501. Available from: https://doi.org/10.1016/j.bone.2007.02.007
- Xu K, Fei W, Gao W, Fan C, Li Y, Hong Y, et al. SOD3 regulates FLT1 to affect bone metabolism by promoting osteogenesis and inhibiting adipogenesis through PI3K/AKT and MAPK pathways. Free Radic Biol Med. 2024;212:65-79. Available from: https://doi.org/10.1016/j.freeradbiomed.2023.12.021
- Park SD, Saunders AS, Reidy MA, Bender DE, Clifton S, Morris KT. A review of granulocyte colony-stimulating factor receptor signaling and regulation with implications for cancer. Front Oncol. 2022;12. Available from: https://doi.org/10.3389/fonc.2022.932608
- Isojima T, Walker EC, Poulton IJ, McGregor NE, Wicks IP, Gooi JH, et al. G-CSF receptor deletion amplifies cortical bone dysfunction in mice with STAT3 hyperactivation in osteocytes. J Bone Miner Res. 2022;37(10):1876-1890. Available from: https://doi.org/10.1002/jbmr.4654
- Semerad CL, Christopher MJ, Liu F, Short B, Simmons PJ, Winkler I, et al. G-CSF potently inhibits osteoblast activity and CXCL12 mRNA expression in the bone marrow. Blood. 2005;106(9):3020-3027. Available from: https://doi.org/10.1182/blood-2004-01-0272
- Katayama Y, Battista M, Kao WM, Hidalgo A, Peired AJ, Thomas SA, et al. Signals from the sympathetic nervous system regulate hematopoietic stem cell egress from bone marrow. Cell. 2006;124(2):407-421. Available from: https://doi.org/10.1016/j.cell.2005.10.041
- Khoswanto C. Role of matrix metalloproteinases in bone regeneration: narrative review. J Oral Biol Craniofac Res. 2023;13(5):539-543. Available from: https://doi.org/10.1016/J.JOBCR.2023.06.002
- Krause C, Guzman A, Knaus P. Noggin. Int J Biochem Cell Biol. 2011;43(4):478-481. Available from: https://doi.org/10.1016/j.biocel.2011.01.007
- Potti TA, Petty EM, Lesperance MM. A comprehensive review of reported heritable noggin-associated syndromes and proposed clinical utility of one broadly inclusive diagnostic term: NOG-related-symphalangism spectrum disorder (NOG-SSD). Hum Mutat. 2011;32(8):877-886. Available from: https://doi.org/10.1002/humu.21515
- Gong Y, Krakow D, Marcelino J, Wilkin D, Chitayat D, Babul-Hirji R, et al. Heterozygous mutations in the gene encoding noggin affect human joint morphogenesis. Nat Genet. 1999;21(3):302-304. Available from: https://doi.org/10.1038/6821
- Masuda S, Namba K, Mutai H, Usui S, Miyanaga Y, Kaneko H, et al. A mutation in the heparin-binding site of noggin as a novel mechanism of proximal symphalangism and conductive hearing loss. Biochem Biophys Res Commun. 2014;447(3):496-502. Available from: https://doi.org/10.1016/j.bbrc.2014.04.015
- Sémonin O, Fontaine K, Daviaud C, Ayuso C, Lucotte G. Identification of three novel mutations of the noggin gene in patients with fibrodysplasia ossificans progressiva. Am J Med Genet. 2001;102(4):314-317. Available from: https://doi.org/10.1002/ajmg.1504
- Song T, Shi J, Guo Q, Lv K, Jiao X, Hu T, et al. Association between NOGGIN and SPRY2 polymorphisms and nonsyndromic cleft lip with or without cleft palate. Am J Med Genet A. 2015;167A(1):137-141. Available from: https://doi.org/10.1002/ajmg.a.36802
- Gutiérrez-Prieto SJ, Torres-López DM, García-Robayo DA, Rey-Cubillos JA, Gómez-Rodríguez M. Clinical and molecular study of the NOG gene in families with mandibular micrognathism. Eur J Dent. 2021;15(4):746-754. Available from: https://doi.org/10.1055/s-0041-1726162
- Gutiérrez SJ, Gómez M, Rey JA, Ochoa M, Gutiérrez SM, Prieto JC. Polymorphisms of the noggin gene and mandibular micrognathia: a first approximation. Acta Odontol Latinoam. 2010;23(1):13-19. Available from: https://pubmed.ncbi.nlm.nih.gov/20645637/
- Lieber CS. Alcohol and the liver: metabolism of alcohol and its role in hepatic and extrahepatic diseases. Mt Sinai J Med. 2000;67(1):84-94. Available from: https://pubmed.ncbi.nlm.nih.gov/10677787/
- Moreno Otero R, Cortés JR. Nutrition and chronic alcohol abuse. Nutr Hosp. 2008;23 Suppl 2:3-7. Available from: https://pubmed.ncbi.nlm.nih.gov/18714405/
- Kini U, Nandeesh BN. Physiology of bone formation, remodeling, and metabolism. In: Radionuclide and hybrid bone imaging. 2012;29-57. Available from: https://www.spinedragon.com/student_material/reading/2017_bone_formation_remodelling.pdf
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.


 Save to Mendeley
Save to Mendeley
